From Sample to Data
With our easy-to-use workflow, you can rapidly go from sample to data
Sample Prep
Hi-C/CiFi/RNA/ATAC-seq
Compatible with cell lines, primary cells, fresh/frozen tissue, and FFPE
Library Prep
Generate High-Quality Sequencing Libraries
Processed with appropriate library preparation workflow
Sequencing
Illumina or PacBio Next Generation Sequencing
overage and sequencing depth vary based on project requirements & platform
Data Analysis
Powerful Analysis Built on Open Source Tools
Bioinformatics tool based on application
Benefits of Multi-Omics Service
- One partner, complete data — integrate 3D genome structure, chromatin accessibility, and gene expression without coordinating across multiple service providers
- Flexible sample compatibility — work with fresh frozen tissue, cell pellets, FFPE, and low-input samples across mammalian, plant, and other organisms
- Built-in quality assurance — integrated QC checkpoints at every stage deliver reliable, reproducible, publication-quality results
- End-to-end support — from multi-omics library prep through sequencing and bioinformatics, all from a single team
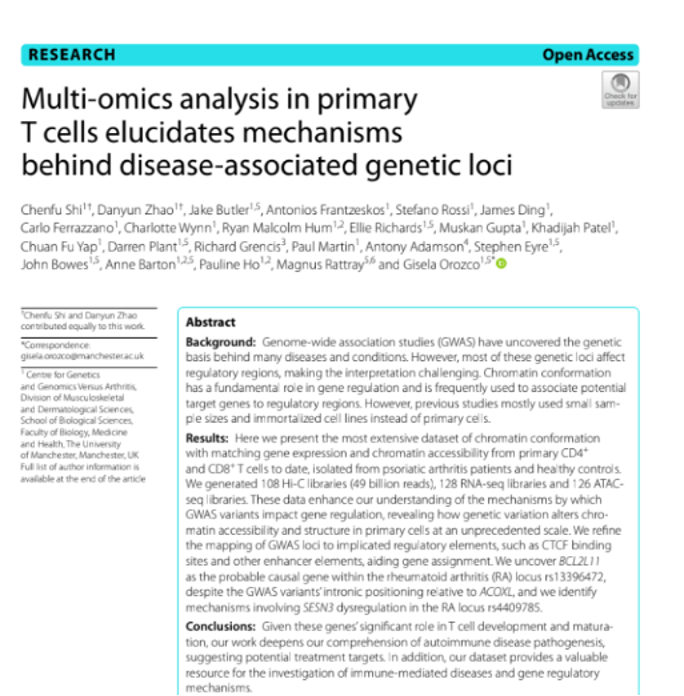
Multi-Omics and 3D Genomics Decode Arthritis Disease Mechanisms
Researchers at the University of Manchester combined Hi-C, RNA-seq, and ATAC-seq data from primary T cells isolated from psoriatic arthritis patients to decode how disease-associated genetic variants influence gene regulation. By integrating 3D chromatin architecture (Hi-C), gene expression (RNA-seq), and chromatin accessibility (ATAC-seq), the team identified specific enhancer-gene connections and TAD boundary disruptions that explain how distant non-coding variants drive autoimmune disease risk.
Shi, C., Zhao, D., Butler, J. et al. Multi-omics analysis in primary T cells elucidates mechanisms behind disease-associated genetic loci. Genome Biol 26, 26 (2025).
Arima Multi-Omics Publications
Explore Our Resources
Learn more about how you can use Arima Hi-C kits and workflows in your research: